For a long time, agricultural fungal disease control has relied heavily on chemical fungicides. However, their overuse has led to environmental pollution, pesticide residues, and the emergence of resistant pathogens—issues that seriously hinder the development of sustainable agriculture. Tapping into plants' natural disease-resistance resources has thus become a core strategy for green control. Over millions of years of evolution, plants have developed a sophisticated innate immune system, in which antifungal peptides (AFPs) serve as key defense components. AFPs typically consist of 2 to 100 amino acids and act through mechanisms such as disrupting fungal cell membranes or inhibiting cell wall synthesis. However, due to their low sequence similarity and short length, traditional bioinformatics tools (such as BLAST) struggle to identify novel AFPs. To unlock the defensive potential hidden in plant genomes, researchers at Shanghai Jiao Tong University developed the first AI model specifically designed for plant AFPs—FungiGuard—which efficiently and accurately identifies antifungal peptides from short open reading frames (sORFs) or non-canonical ORFs, offering a powerful new solution to this challenge.
Building the "Ultimate Brain": Multi-Model Ensemble and Performance Evaluation
Researchers first curated a high-confidence training set from the PlantPepDB database, consisting of 506 plant AFPs and 2,638 non-AFP peptides. They evaluated various machine learning models, including Long Short-Term Memory networks (LSTM), bidirectional LSTM, attention-enhanced variants, and Random Forest Classifier (RFC). On an independent test set, RFC performed the best (AUC = 0.93). By implementing a five-model majority voting ensemble, the FungiGuard framework boosted prediction precision to 89.58%, significantly outperforming existing general-purpose AFP predictors like AFP-MFL.
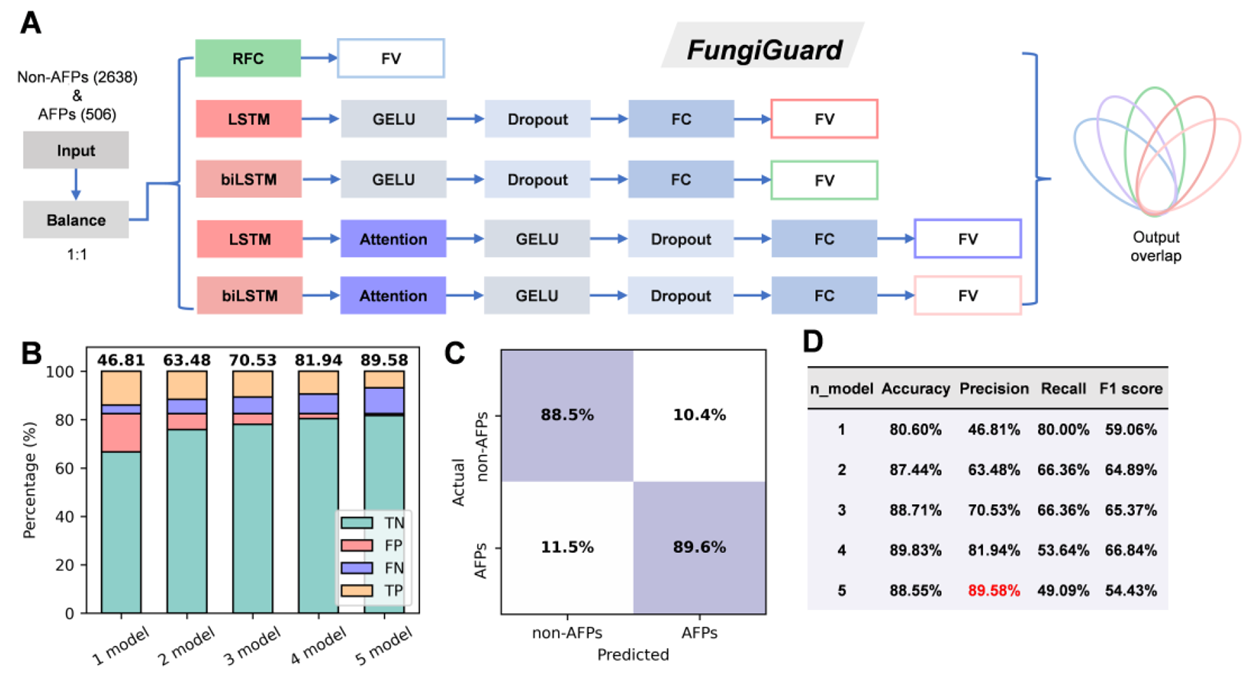
Figure 1. FungiGuard architecture and performance in plant AFP classification
Plant-Specific Optimization Delivers Superior Performance
When researchers compared FungiGuard head-to-head with the leading general-purpose antibacterial peptide predictor AFP-MFL, they found that AFP-MFL had a false positive rate as high as 66.5% on plant-derived peptides, whereas FungiGuard achieved more than three times higher precision on the same test set. This result highlights the critical need for domain-specific optimization tailored to plants, establishing FungiGuard as the most suitable AFP identification tool for plant science research today.
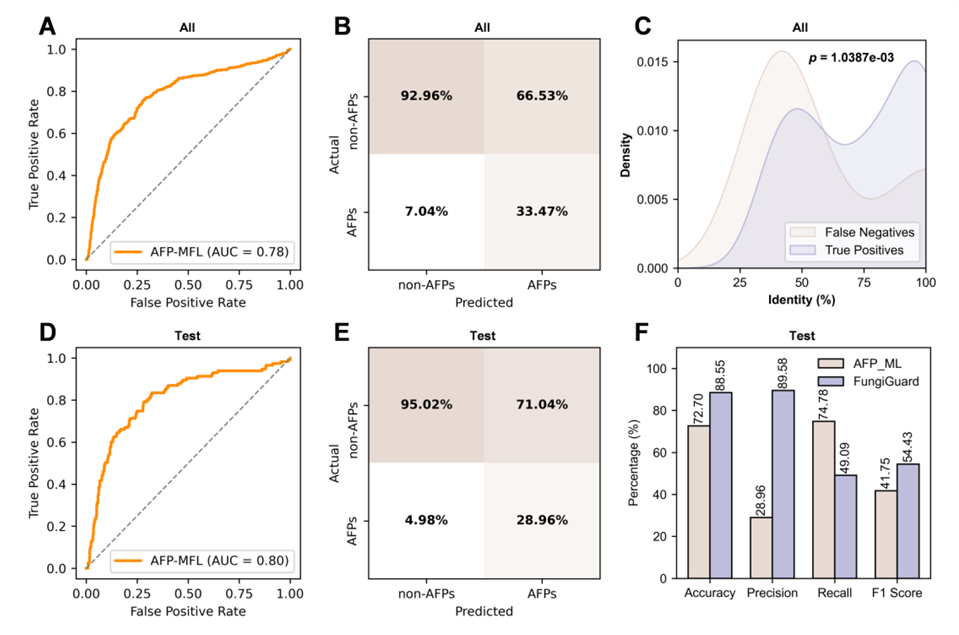
Figure 2. Performance comparison between AFP-MFL and FungiGuard
Systematic Screening of Novel Defense Molecules in the Arabidopsis Genome
Using FungiGuard to screen short open reading frames (sORFs) in Arabidopsis, wheat, rice, and maize, the team identified 35 high-confidence AFP candidates in Arabidopsis. These candidates were significantly enriched in biological processes such as "defense response to fungi." Transcriptomic analysis further confirmed that genes like AT5G44430—previously not annotated as defense peptides—showed strong upregulation upon fungal infection, revealing a wealth of previously hidden functional peptides in the genome.
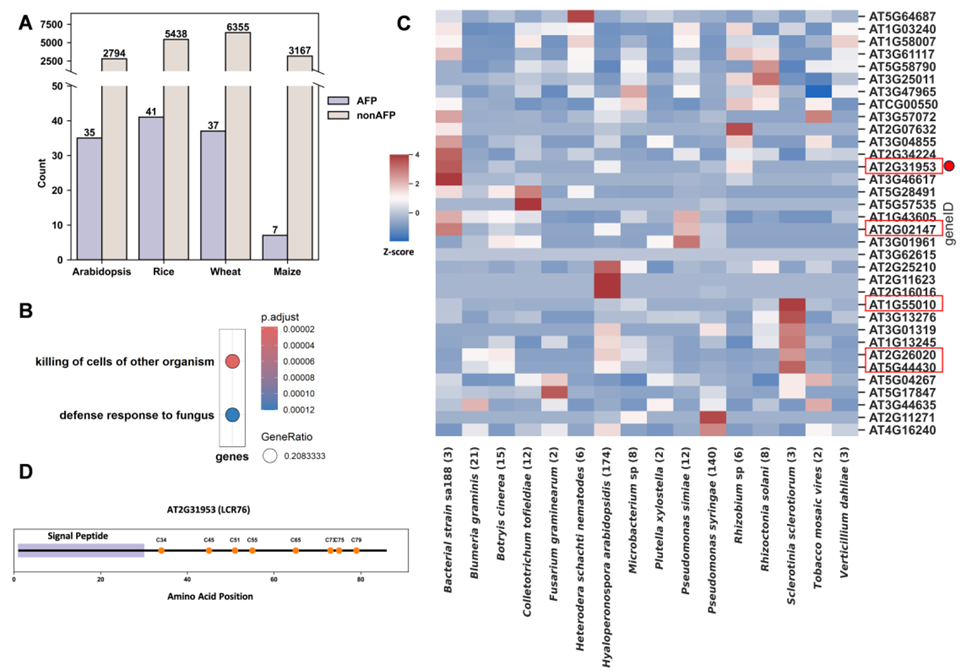
Figure 3. Identification of AFPs from short open reading frames in four plant species using FungiGuard
Mining Disease-Resistance Newcomers from Non-Coding Regions: AtcAFP5
Going further, the researchers applied the model to scan genomic regions previously considered non-coding (such as uORFs and dORFs), successfully identifying 11 non-canonical peptides (NCPs) induced by Botrytis cinerea, named AtcAFP1–11. Through in vitro assays and in vivo spray experiments, they confirmed that one of them, AtcAFP5, exhibits excellent antifungal activity, with a minimum inhibitory concentration (MIC) of 16 μmol/L and a significant reduction in lesion area on infected leaves.
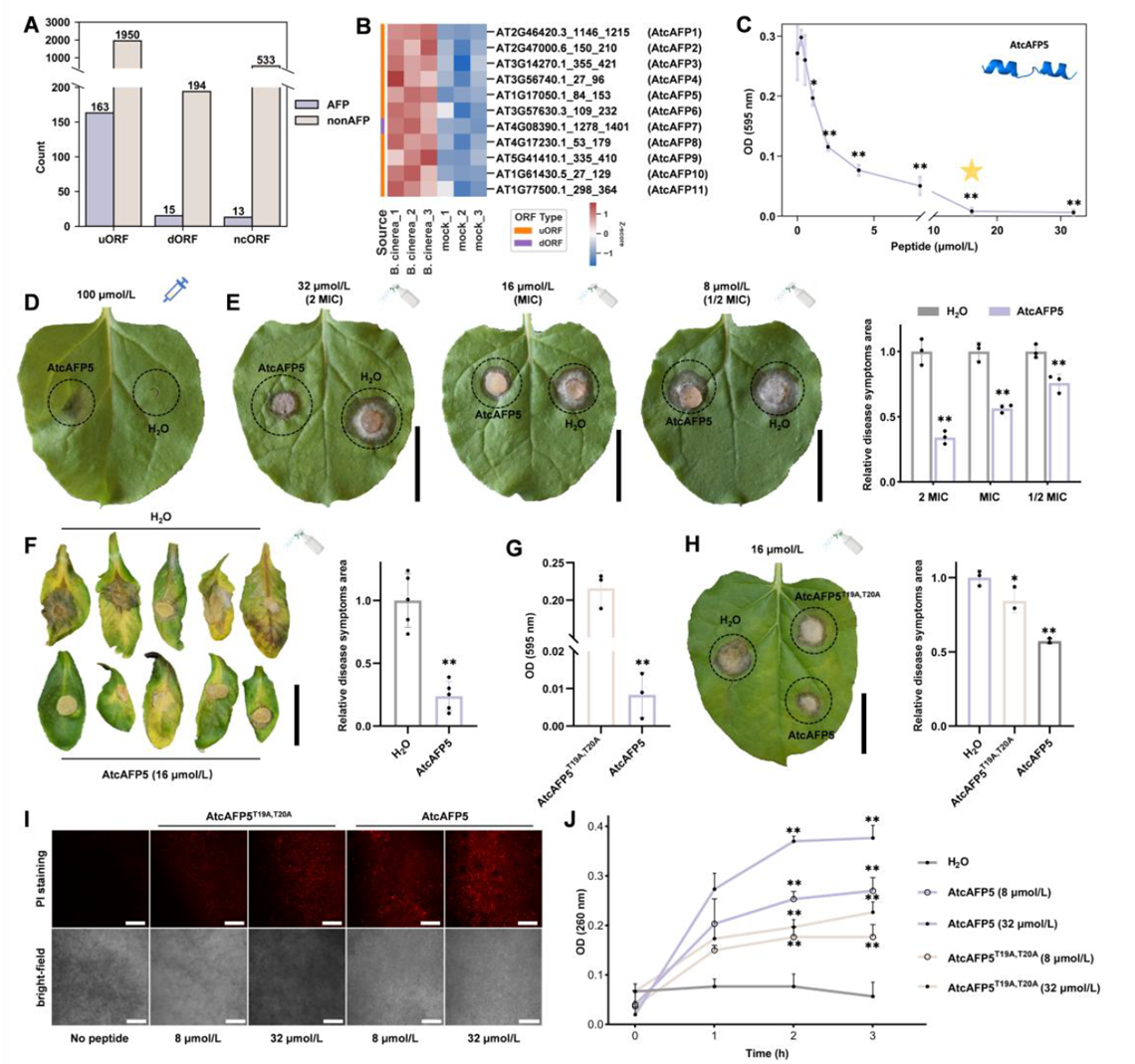
Figure 4. Identification and functional validation of Arabidopsis non-canonical peptides (NCPs) with activity against Botrytis cinerea (gray mold)
Expanding Chemical Space with Random Sequence Generation
To explore AFP possibilities beyond nature, the team generated 10,000 random peptides mimicking the amino acid preferences of natural AFPs (high cysteine and glycine content). FungiGuard screened out 514 potential high-activity candidates. Statistical analysis showed that these predicted AFPs closely matched known AFPs in secondary structure (high α-helix content) and physicochemical properties such as isoelectric point—validating the model's strong generalization ability and providing design templates for synthetic antifungal peptides.
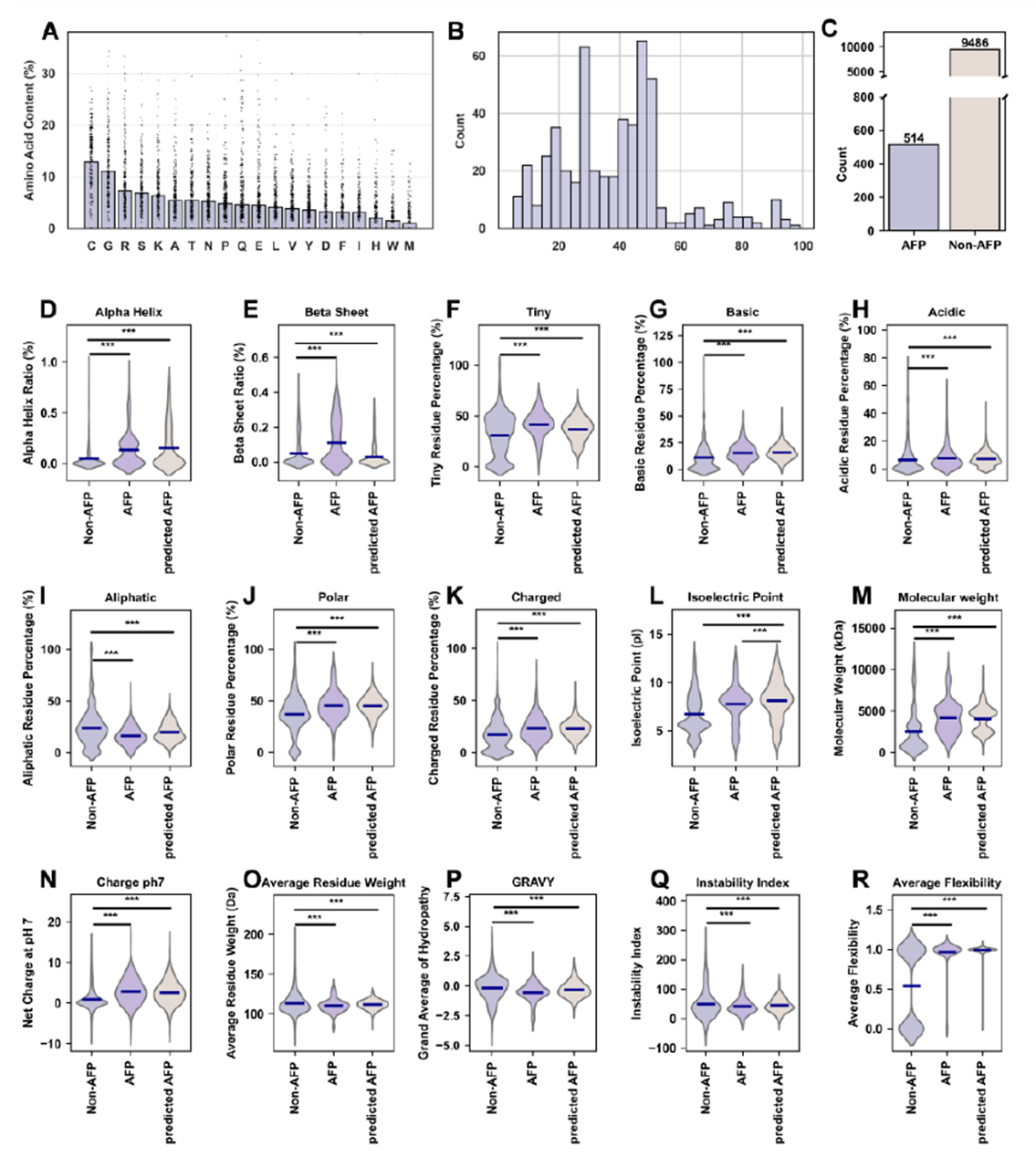
Figure 5. Comparison of features between FungiGuard-predicted AFPs and known AFPs in the plant small peptide dataset
Study Limitations and Future Directions
The study also acknowledges limitations: the training set has limited species diversity among plant peptides, introducing some taxonomic bias; experimental validation focused only on Botrytis cinerea and has not yet explored broad-spectrum antifungal activity of the candidates. Future work could expand the training dataset, incorporate advanced models such as BERT or Mamba for further optimization of FungiGuard, conduct activity testing against multiple fungal pathogens, and investigate the immune-modulatory mechanisms of multifunctional AFPs—laying a stronger foundation for their practical application in agricultural production.
abinScience Experimental Support
abinScience provided the anti-GFP antibody for this study. Experimental application: Researchers used this antibody in Western Blot experiments to verify the stable expression of AtcAFP5-GFP fusion protein in plant cells. This helped confirm correct expression and size of the fusion protein, providing key experimental evidence for subsequent confocal microscopy observation of AtcAFP5 localization in fungal and plant cells, as well as elucidation of its mechanism in disrupting fungal cell membranes.
For research use only. Not for use in diagnostic or therapeutic procedures.

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский







