As key effectors of innate immunity, NK cells play a pivotal role by directly recognizing and eliminating abnormal cells without prior sensitization. Yet their marked heterogeneity and multi-faceted functionality make precise characterization a formidable challenge.OMIP-007, a 14-color flow cytometry panel, delivers robust technical support for in-depth investigations into NK cell phenotype, differentiation, activation and migration.
By systematically incorporating key markers across four critical dimensions—NK cell identity, differentiation, function and migration,OMIP-007 serves as a standardized tool for dissecting NK cell roles in anti-infective defense, tumor suppression and immune regulation. It empowers researchers to delve into the immune properties of NK cells across a wide range of disease settings.
OMIP-007 not only acts as a precision tool for targeted research areas like HIV infection, but also offers a versatile reference framework for immunological studies spanning multiple diseases. Its sound design principles and transparent data output lay a robust foundation for future research endeavors in this field.
Natural Killer (NK) cells, as a core component of innate immunity, can rapidly recognize and eliminate abnormal cells without prior sensitization. Fine phenotypic analysis of NK cells is crucial to understanding their roles in anti-infection, anti-tumor, and immune regulation. OMIP-007, through a 14-color panel, is the first to systematically integrate key markers across the four dimensions of NK cell identity, differentiation, function, and migration in a multi-color flow cytometry system. This panel not only provides precise tools for HIV infection research but also serves as a standardised reference for in-depth exploration of NK cell immune characteristics in a variety of disease contexts.
| Target | Fluorochrome | Function | abinScience Recommendation |
|---|---|---|---|
| Live/Dead | AqBlu | Exclude dead cells, T cells, B cells, monocytes | — |
| CD3 | PE-TR | View CD3 antibodies | |
| CD4 | ECD | View CD4 antibodies | |
| CD14 | PE-Cy5 | View CD14 antibodies | |
| CD19 | PE-Cy5 | View CD19 antibodies | |
| CD16 | Pac Blue | NK cells | View CD16 antibodies |
| CD56 | PE-Cy7 | View CD56 antibodies | |
| CD8 | APC-H7 | View CD8 antibodies | |
| α4β7 | Qdot 655 | Homing | View α4β7 antibodies |
| CD62L | Qdot 605 | View CD62L antibodies | |
| KIR2DL1/DS1 | PerCP-Cy5.5 | KIR receptors | View KIR antibodies |
| KIR2DL2/DS2/DL3 | PE | ||
| KIR3DL1 | Alexa 700 | ||
| HLA-DR | FITC | Activation/ Differentiation | View HLA-DR antibodies |
| CD57 | APC | — |
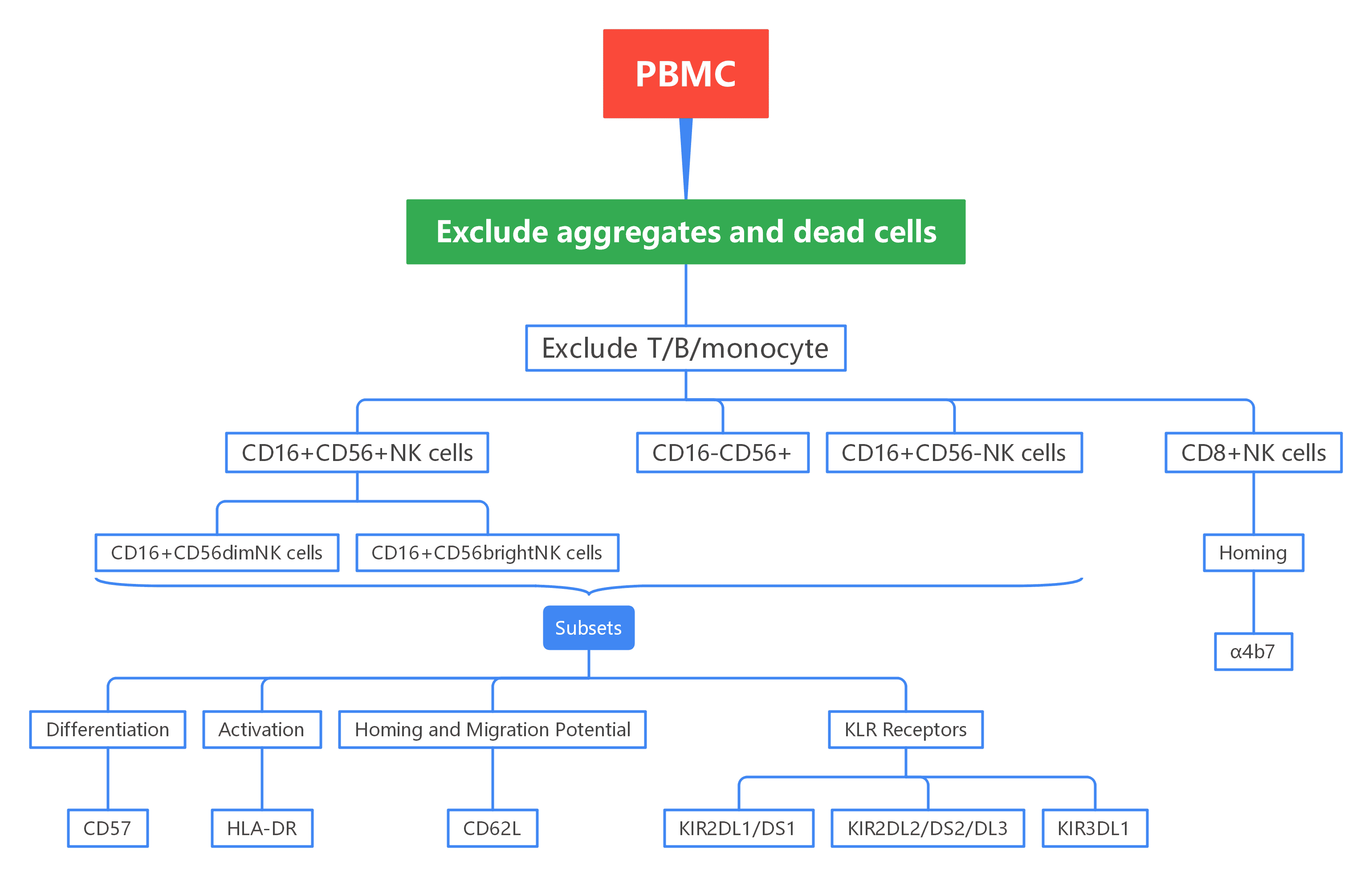
Figure 1. Overview of OMIP-007 Gating Strategy
Cell aggregates and dead cells were removed to ensure that only viable cells were used for analysis.
T cells, monocytes, and B cells were excluded using specific markers. NK cells were then identified by CD56 and CD16, distinguishing CD56hi, CD56dim, and CD56- subsets.
Markers like CD57, CD62L, and HLA-DR were used to assess NK cell activation, maturation, and homing capabilities.
KIR receptor co-expression patterns were analyzed using three KIR antibodies, revealing NK cell maturation and function.
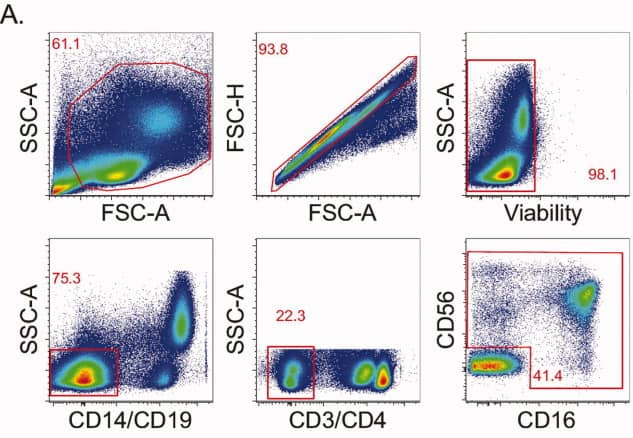
1). NK-cell identification
Lymphocytes were first gated by FSC/SSC, exclude aggregates and non-viable cells. CD14, CD19, CD3, and CD4 were used to exclude monocytes, B cells, and T cells. NK cells were subsequently identified and stratified into major subsets on the basis of CD56 and CD16 expression.
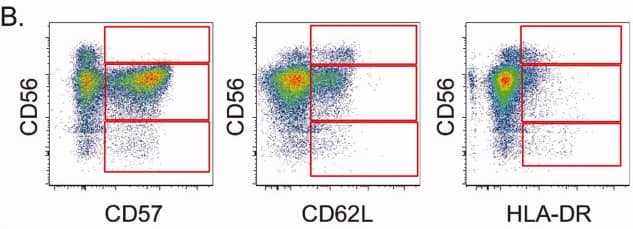
2). Differential expression of CD57, CD62L, and HLA-DR
Across CD56hi, CD56dim, and CD56- NK-cell subsets, the panel reveals distinct expression patterns of CD57 (maturation marker), CD62L (lymphoid-homing molecule), and HLA-DR (activation marker), providing a multi-layered view of NK-cell differentiation and activation states.
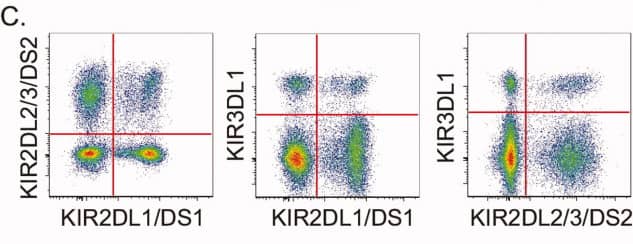
3). KIR receptor combinations
Differentiate CD4+ and CD8+ T cell phenotypes using CD45RA, CD28, and CCR7 markers (grey markers), and project cytokine-secreting cells (blue) from panel B onto panel C to observe the cytokine secretion capabilities of different cell types.
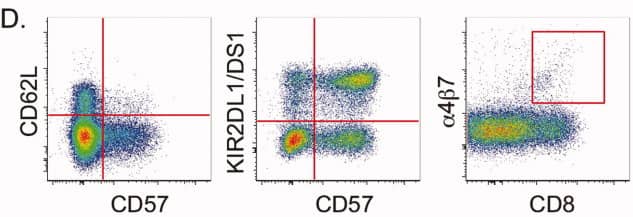
4). Key phenotypic relationships
CD62L and CD57
The panel demonstrates that lymphoid-homing potential (CD62L) and terminal differentiation (CD57) are largely mutually exclusive within NK-cell populations.
KIR2DL1/DS1 and CD57
The coordinated acquisition of KIR receptors together with CD57 expression reflects NK-cell maturation trajectories.
α4β7 and CD8
A distinct subset of CD8+ NK cells expressing the gut-homing receptor α4β7 is identified, emphasising the diversity of NK-cell migratory phenotypes.
4.1 A logically structured panel covering the essential dimensions of NK biology
This 14-color panel enables multi-dimensional analysis of NK cell phenotype, differentiation, activation, and migration by integrating key surface markers. It distinguishes NK cell subsets (CD56hi, CD56dim, and CD56-) using CD56 and CD16, combines KIR receptors with CD57 to define differentiation and maturation pathways, and assesses activation and tissue migration potential through markers such as HLA-DR, CD62L, and α4β7.
4.2 Introducing multi-KIR co-expression analysis into a single-tube phenotyping panel
OMIP-007 combines several KIR antibodies into a single validated panel, allowing for the identification of co-expression patterns of KIR receptors at the single-cell level. This approach provides a foundation for linking phenotypic data with individual genotypes or functional assessments in future studies.
4.3 Transparent rationale and reporting of excluded markers—valuable experimental guidance
OMIP-007 provides detailed data on the testing and exclusion of key markers like CCR7, CD38, and NKG2A. For example, it clarifies that CD38 is constitutively expressed at high levels on NK cells, making it unsuitable as an activation marker, while CCR7 expression is too low for detection. This transparency in reporting negative results offers valuable guidance to avoid common experimental pitfalls, preventing wasted resources, and enhancing the panel's credibility as a reliable tool.
OMIP-007 establishes a comprehensive phenotypic analysis system for human NK cells using a 14-color panel, optimized for cryopreserved PBMCs. It enables simultaneous assessment of NK cell subset distribution, maturation status, and activation/migration profiles, while also facilitating the identification of co-expression patterns of multiple KIR receptors in a single-tube assay—laying a solid foundation for in-depth exploration of their specific functions in immune responses.
This panel strikes an ideal balance between clinical sample compatibility and the depth of phenotypic characterization, delivering a standardized and reliable technical framework for dissecting NK cell biology across diverse immunological research contexts, including infection and oncology studies.
abinScience provides validated Flow Cytometry Antibodies covering key targets in this panel, supporting your NK cell research
[1] Foulds KE, Donaldson M, Roederer M. OMIP-005: Quality and phenotype of antigen-responsive rhesus macaque T cells. Cytometry A. 2012;81(5):360-361.

As a strategic venture of AtaGenix (established 2011), abinScience was founded in 2023 to deliver premium life science reagents that accelerate discovery. Our flow cytometry antibody products cover commonly used detection markers, with a wide variety to meet the research needs of multiple species (Human/Mouse/Rat/Dog/Hamster/Monkey, etc.). We provide stable and reliable support for scientific research.
+86-027-65523339
중국 우한시 심둔사로 666번지 C동, 우한, 430206

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский