The Complexity of T-Cell Differentiation Profiling
T cells undergo a complex differentiation process during immune responses, forming functionally diverse subsets. Their dynamic changes directly impact immune memory formation, vaccine responses, and the efficacy of immunotherapies. Traditional analytical methods often fail to fully capture this continuous and multi-layered differentiation trajectory, limiting deeper insights into their functional states.
Systematic Mapping of T-Cell Differentiation Dynamics
OMIP-013 integrates key markers such as CD95 and CD127 to advance T-cell analysis from static classification to a logically structured developmental pathway. This approach not only distinguishes classical memory subsets but also tracks the complete differentiation continuum from stem cell-like memory T cells (TSCM) to terminal effector cells, offering a visual tool for studying immune cell evolution.
Simultaneous Analysis of Rare Subsets and Functional States
Beyond covering major T-cell populations, OMIP-013 places special focus on rare subsets such as recent thymic emigrants (RTE) and incorporates functional markers like CD244 to concurrently assess inhibitory states. This integrated design enables the acquisition of differentiation, origin, and functional information in a single experiment, significantly enhancing data dimensionality and experimental efficiency.
Developed by Mahnke et al. (2012), OMIP-013 is a multicolor flow panel designed to study human CD4+/CD8+ T-cell differentiation. By combining 13 markers, it establishes a coherent analytical framework that enables systematic dissection of differentiation pathways alongside assessment of rare subsets and functional states, offering a reliable standardized tool for immunological research.
1. OMIP-013 Panel
2. Gating Logic
1
Live T-Cell Isolation
PBMCs, exclude aggregates and dead cells, gate on CD3+ T cells, and separate into CD4+ and CD8+ subsets.
2
Memory Subset Identification
Within CD4+/CD8+ T cells, perform initial classification based on CCR7 and CD45RA expression, followed by functional refinement using CD27.
3
Rare Precursor Identification
Among naïve phenotype cells, further distinguish true naïve T cells from RTE using CD95 and CD31; identify TSCM within specific populations using additional marker combinations.
4
Functional State Correlation
Analyze the expression of functional markers (e.g., CD244) within the defined/gated subsets.
3. Experimental Results
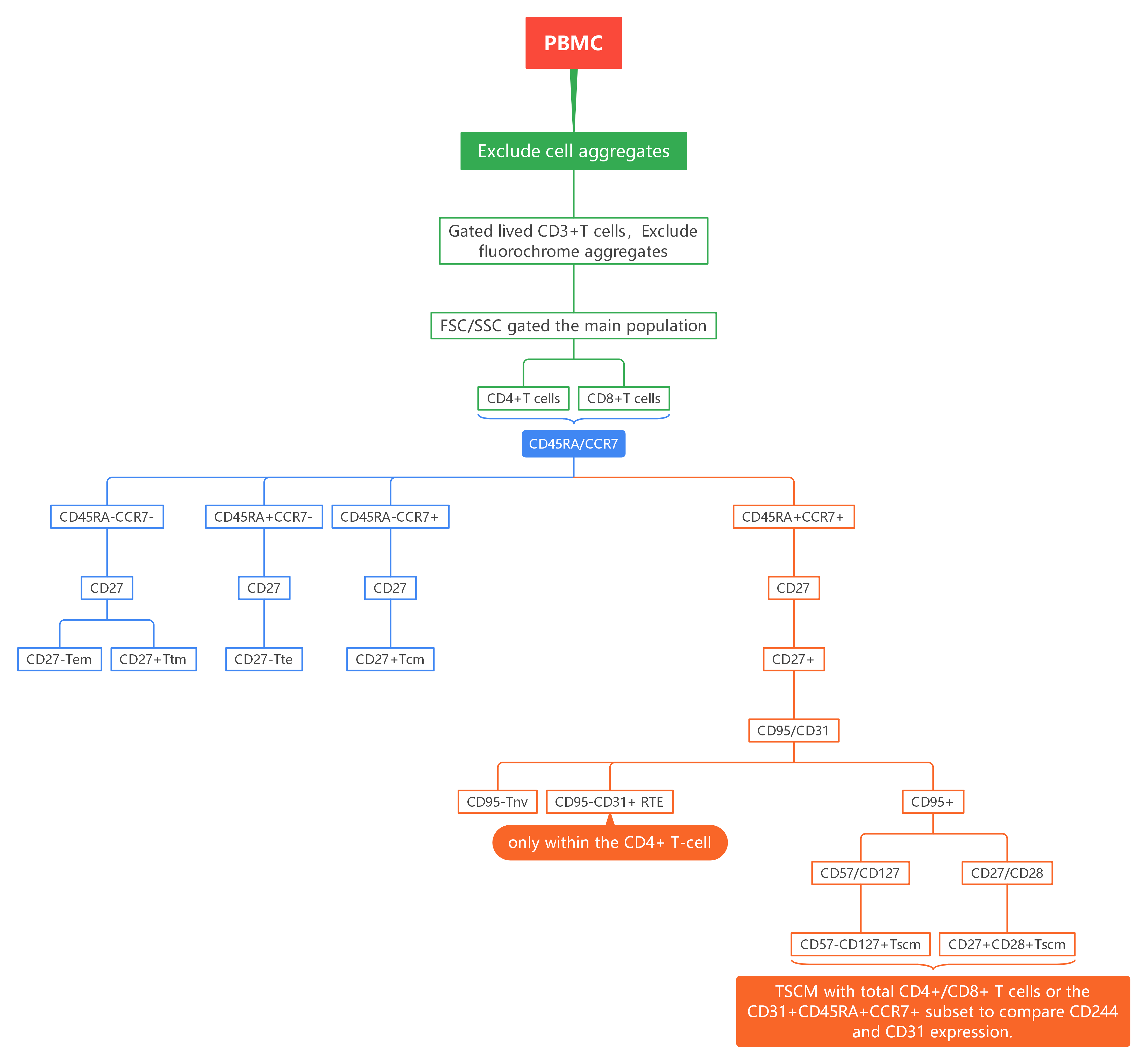
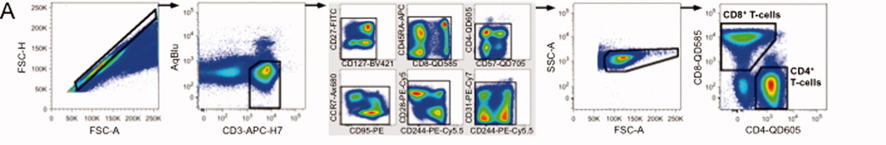
1). PBMCs, exclude aggregates, gate on live CD3+ T cells, exclude fluorochrome aggregates, select the main cell population, and separate into CD4+ and CD8+ T cells.
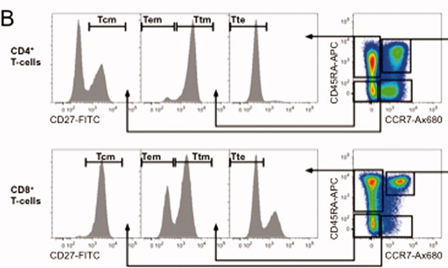
2). Within CD4+ and CD8+ T cells, distinguish activation subsets using CCR7/CD45RA, then further delineate using CD27: Tem (CD45RA-CCR7-CD27-), Ttm (CD45RA-CCR7-CD27+), Tte (CD45RA+CCR7-CD27-), Tcm (CD45RA-CCR7+CD27+).
3). Among CD45RA+CCR7+CD27+ cells, use CD95/CD31 to distinguish: Tnv (CD95-), RTE (CD95-CD31+). Within CD95+ cells, gate CD57-CD127+ TSCM; TSCM can also be identified via CD27/CD28 (CD27+CD28+). Finally, overlay the gated TSCM with total CD4+/CD8+ T cells or the CD31+CD45RA+CCR7+ subset to compare CD244 and CD31 expression.
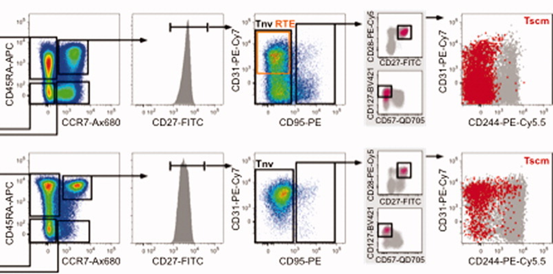
*Note: RTE cells are present only within the CD4+ T-cell compartment.
4. Panel Interpretation
4.1 Systematic Definition of T-Cell Differentiation Pathways
OMIP-013 introduces CD95 (Fas) and CD127 (IL-7Rα) to transform differentiation analysis from a simple classification scheme into a dynamic trajectory with developmental logic and functional directionality. This allows researchers to map the progressive pathway from TSCM → TCM → TEM → TTE and to isolate the therapeutically promising TSCM population at the starting point. This is highly valuable for understanding immune memory formation, vaccine response mechanisms, and quality assessment of cellular therapy products like CAR-T cells.
4.2 Integration of Rare Subsets and Functional States
(1) Identification of recent thymic emigrants (RTE): Among CD4+ naïve T cells (CD45RA+CCR7+), RTE are defined by CD31 (PECAM-1) expression, enabling simultaneous evaluation of thymic output in routine differentiation analysis.
(2) Incorporation of functional marker CD244: After clear delineation of all major and rare subsets, CD244 expression is analyzed within each subset. As a co-inhibitory receptor associated with T-cell exhaustion, its expression provides direct insight into the functional state of T cells at different differentiation stages.
4.3 Highly Structured Experimental Design Philosophy
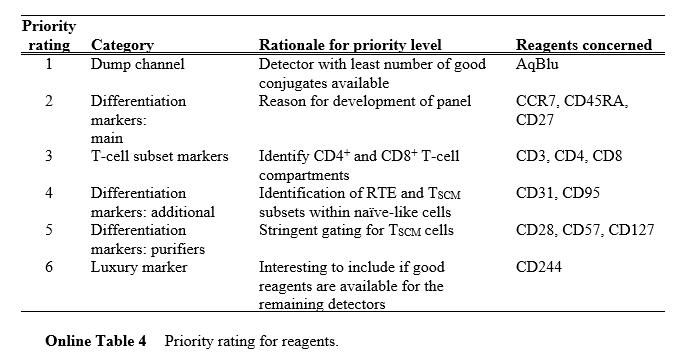
During development, OMIP-013 employed a priority-tiered strategy (Online Table 4), categorizing antibodies into "Essential," "Supplemental," and "Optional" tiers. This ensures detection of core markers is prioritized within limited channels, offering a methodological reference for multicolor panel design.
5. Applications
Immune Memory Studies Vaccine Efficacy Evaluation Cell Therapy Product QC (e.g., CAR-T) T-Cell Dynamics in Autoimmunity Infection & T-Cell Exhaustion Immunosenescence & Thymic Function Tumor-Infiltrating T-Cell Analysis Immune Reconstitution Monitoring
6. Conclusion
Through innovative panel design and optimized reagent combinations, OMIP-013 enables efficient identification of T-cell differentiation states. Its major strength lies in the detailed analysis of various T-cell differentiation stages, particularly the precise identification of subsets like TSCM and RTE, granting it broad applicability in immunological research.
Get OMIP-013 Compatible Flow Cytometry Antibodies
abinScience provides validated flow cytometry antibodies covering key targets in this panel, supporting your T-Cell Differentiation research
References
[1] Mahnke YD, Beddall MH, Roederer M. OMIP-013: differentiation of human T-cells. Cytometry A. 2012 Nov;81(11):935-6.

About Us
As a strategic venture of AtaGenix (established 2011), abinScience was founded in 2023 to deliver premium life science reagents that accelerate discovery. Our flow cytometry antibody products cover commonly used detection markers, with a wide variety to meet the research needs of multiple species (Human/Mouse/Rat/Dog/Hamster/Monkey, etc.). We provide stable and reliable support for scientific research.
Explore abinScience Flow Cytometry Antibodies

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский