In immunological research, antigen-specific T cells are crucial for understanding antiviral responses and vaccine efficacy. However, these cells are often present at very low frequencies and exhibit distinct differentiation profiles depending on the pathogen involved. OMIP-002 integrates pMHC-I multimer technology with multicolour flow cytometric phenotypic analysis, enabling the simultaneous detection of multiple antigen-specific CD8+ T cells in a single experiment. This approach maps these rare populations onto differentiation trajectories, revealing unique patterns of immune responses specific to different viruses.
1. OMIP-002 Panel
|
Target |
Fluorochrome |
Function |
|
CD3 |
APC-Cy7 |
T cell lineage |
|
CD4 |
QD705 |
|
|
CD8 |
PE- Alexa 594 |
|
|
CD14 |
Pacific Blue |
Exclude dead cells, B cells, monocytes, macrophages |
|
CD19 |
Pacific Blue |
|
|
Dead cells |
VIVID |
|
|
CD27 |
PE-Cy5 |
Memory cells, differentiation |
|
CD127 |
PE- Alexa 700 |
|
|
CCR7 |
PE- Alexa 750 |
|
|
CD45RO |
APC- Alexa 700 |
|
|
CD57 |
FITC |
|
|
PD-1 |
Biotin-APC |
|
|
Multimer 1 |
PE |
Different pMHC-1 polypeptide polymers |
|
Multimer 2 |
QD565 |
|
|
Multimer 3 |
QD605 |
|
|
Multimer 4 |
QD800 |
2. Gate Logic
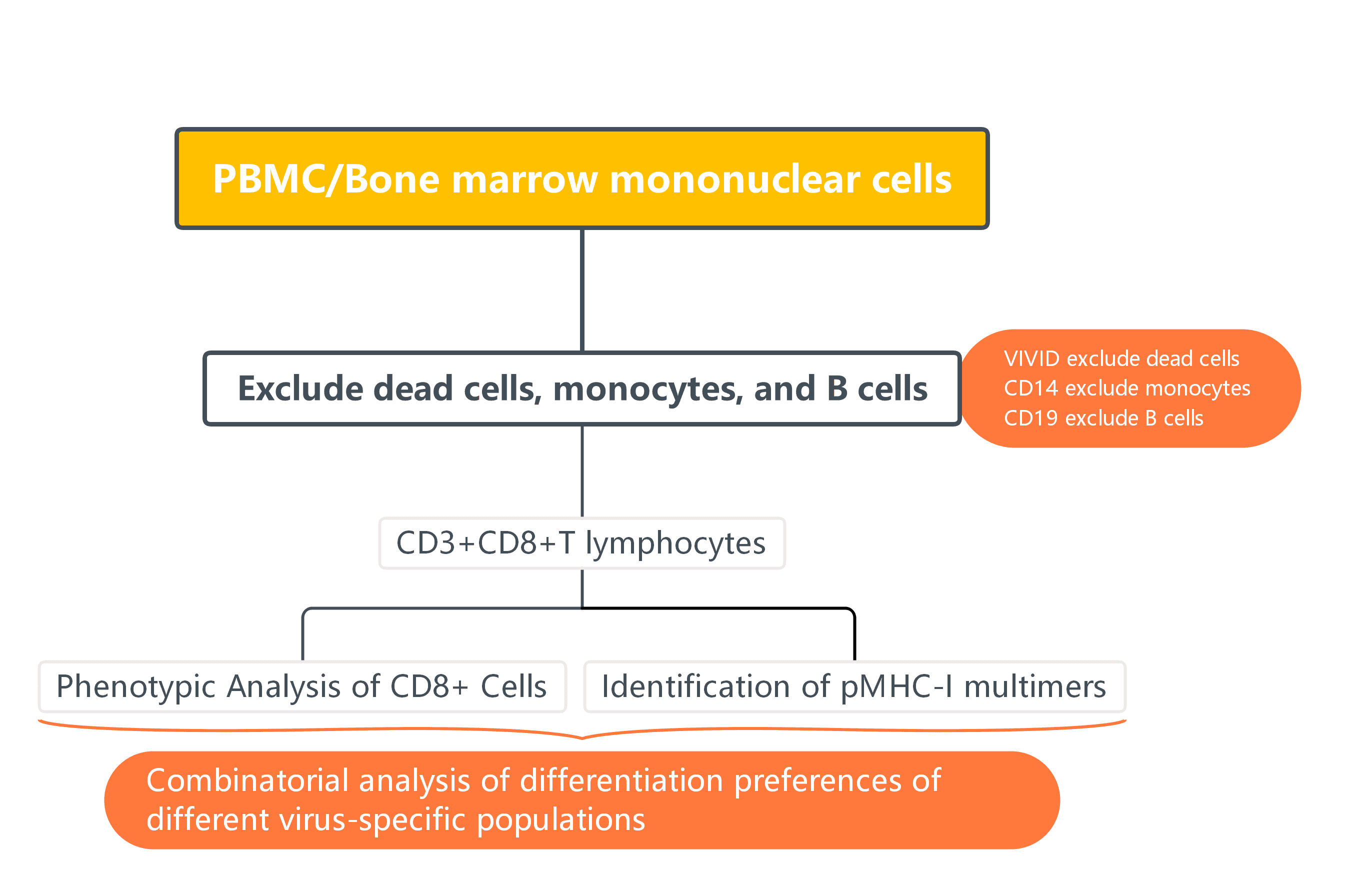
3. Experimental Results
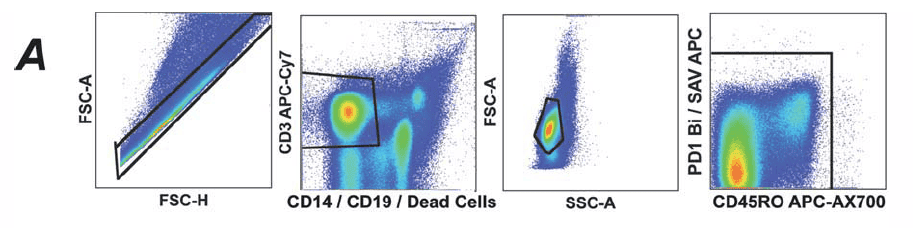
1). FSC-A/H are used to exclude aggregates, then lived CD3+ cells are gated by combining CD3+CD14-CD19-VIVID. Further, non-specific signals caused by dye aggregates are excluded using combinations of markers.
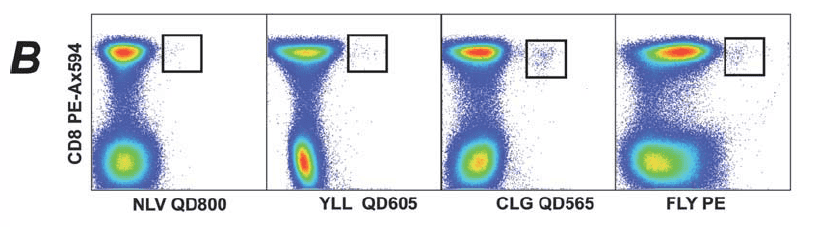
2). Identification of pMHC-I multimers binding to CD8+ T cells: pMHC-I multimers are used to identify cells that recognise pMHC-I peptide complexes within the CD3+ population.
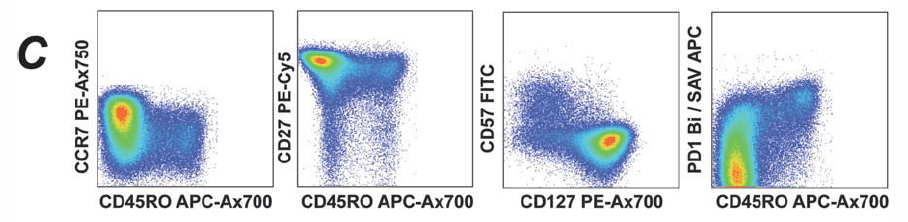
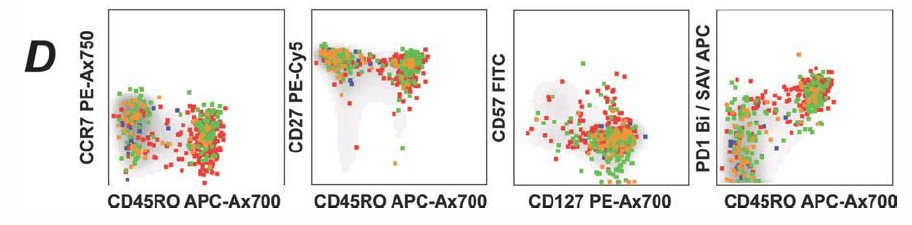
3). Phenotypic Analysis of CD8+ Cells: Further classification of different cell phenotypes within the CD8+ population.
4). Projection of Multimer+ Events: Multimer+ events from panel B are projected onto panel C to illustrate the expression of different pMHC-I peptide multimers on various phenotypic cell subsets.
4. Protocol Interpretation
1). Parallel Detection of Multiple Antigen-Specific Cells:
In a single experiment, four pMHC-I multimers capture CD8+ T cells specific to different epitopes of CMV and EBV. This approach resolves the low throughput issue for detecting rare populations, significantly increasing experimental efficiency.
2). Direct Mapping of Antigen-Specific Populations to Phenotypes:
The multimer+ cells are mapped to differentiation markers such as CCR7, CD45RO, and CD27, providing a clear view of the differentiation preferences of various virus-specific T cell populations. For the first time, the identification and differentiation phenotypes of rare antigen-specific T cells have been comprehensively integrated into a single panel.
3). Standardised and Reproducible Panel Design:
The marker layout and fluorochrome selection in the panel have been optimised to clearly distinguish low-frequency multimer signals and differentiation phenotypes. This provides a reproducible experimental template for multicolour flow cytometry.
5. Conclusion
OMIP-002 showcases an accurate flow cytometry strategy for capturing rare antigen-specific T cells and directly analysing their differentiation states. This method not only answers the question of "how many" but also reveals "what state" the cells are in, making it a classic approach for studying viral responses, vaccine immunology, and immunosenescence.
References:
[1] Chattopadhyay, P.K., Roederer, M. and Price, D.A. (2010), OMIP-002: Phenotypic analysis of specific human CD8+ T-cells using peptide-MHC class I multimers for any of four epitopes†. Cytometry, 77A: 821-822.
+86-027-65523339
중국 우한시 심둔사로 666번지 C동, 우한, 430206

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский