Colorectal cancer (CRC) ranks among the leading causes of cancer incidence and mortality worldwide, with its disease burden continuously increasing. The pathogenesis of CRC is a complex process involving multiple genes and signalling pathways. Mutations in key driver genes (such as APC and KRAS) trigger dysregulation of essential pathways, accompanied by epigenetic alterations and remodelling of the tumour microenvironment, ultimately promoting the transformation of normal colorectal cells into malignancies.
1. Colorectal Cancer Signalling Pathways
1.1 Wnt/β-catenin Signalling Pathway
The Wnt/β-catenin pathway is a core oncogenic pathway in colorectal cancer, with abnormalities observed in approximately 80% of CRC patients. Under normal conditions, β-catenin is degraded via a phosphorylation pathway mediated by GSK-3β, maintaining pathway stability. Mutations in the APC gene or loss of Axin function result in the accumulation of β-catenin in the cytoplasm, which translocates to the nucleus and binds to Tcf/Lef transcription factors, activating proliferation-associated genes like MYC and Cyclin D1. This event is considered a hallmark of early colorectal carcinogenesis.
1.2 EGFR Signalling Pathway
The epidermal growth factor receptor (EGFR) is a transmembrane receptor that activates downstream Ras-Raf-MEK-ERK and PI3K-Akt-mTOR pathways upon ligand binding, regulating cell proliferation, migration, and survival. In CRC, approximately 60% of cases exhibit overexpression of EGFR, and KRAS mutations can lead to ligand-independent, sustained activation of this pathway, offering a critical entry point for EGFR regulatory network studies.
1.3 PI3K/Akt/mTOR Signalling Pathway
The PI3K/Akt/mTOR pathway is abnormally activated through mutations in the PIK3CA gene or loss of the PTEN gene, contributing to tumour cell metabolic reprogramming and the development of drug resistance. Key molecules within this pathway, such as Akt and mTOR, are closely linked to CRC cells' ability to adapt to external stressors, positioning them as pivotal targets for research into tumour cell survival mechanisms and combination therapy strategies.
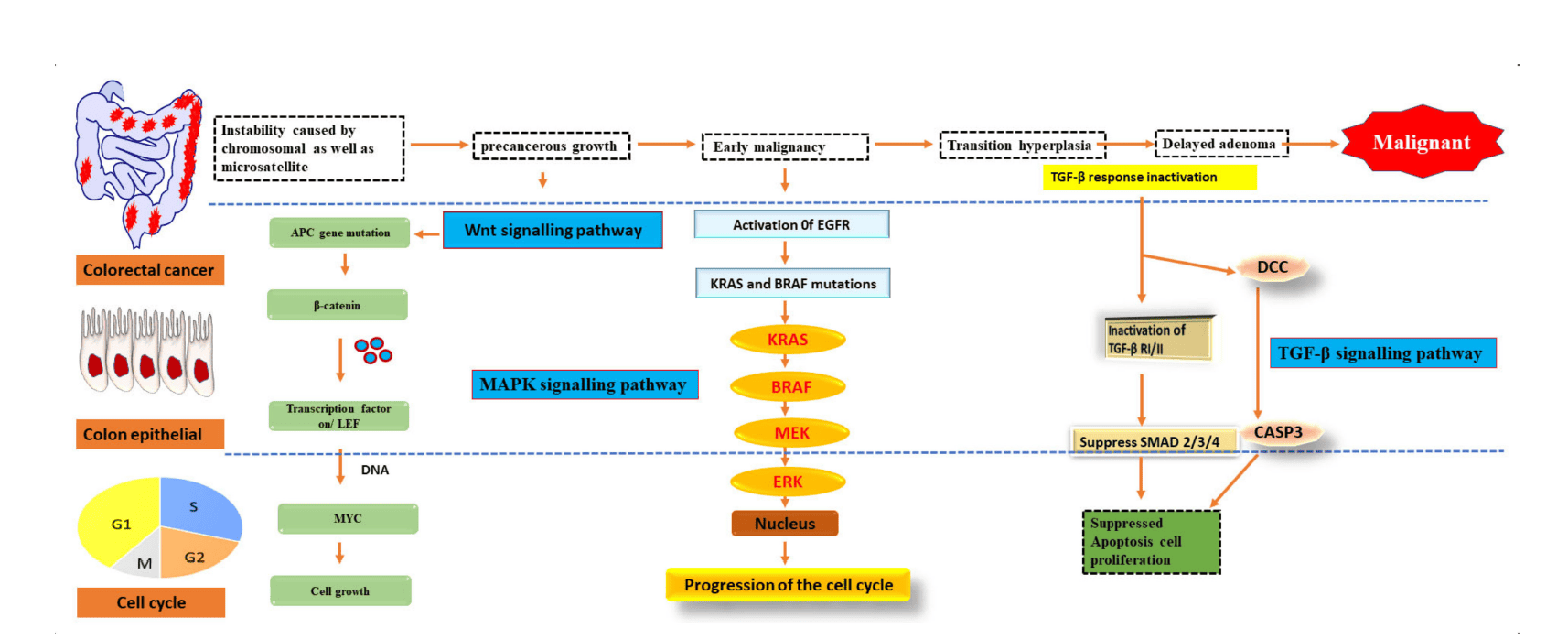
Figure 1: Signalling pathways genetically altered in CRC.
2. Current Research Landscape
2.1 Targeted Regulation Mechanisms
Current research is focused on targeted regulation of classic driver genes in CRC, such as the development of small molecule inhibitors targeting KRAS mutations, particularly the G12C variant. These studies explore how these inhibitors bind KRAS and block downstream pathways. Additionally, collaborative regulatory research on the BRAF mutation-related pathway is ongoing, aiming to clarify the network effects of multi-target combination interventions.
2.2 Immune-Related Mechanisms
Studies investigating the immune microenvironment of CRC are progressing, with a focus on the regulatory expression of PD-1/PD-L1 molecules and the relationship between the MSI-H/dMMR phenotype and immune responses. Research has also revealed that mutations in POLE/POLD1 affect tumour immunogenicity, providing new perspectives on the immune evasion mechanisms of CRC.
2.3 Early Molecular Mechanisms Exploration
Research in early detection primarily targets molecular biomarkers for early-stage colorectal cancer, such as the analysis of KRAS mutations and BMP3/NDRG4 methylation in stool samples. These markers are explored for their potential correlation with colorectal epithelial cell malignancy, with findings validated using cell and animal models to understand the molecular changes during the early stages of CRC.
3. Hot Research Targets in Colorectal Cancer
|
Target |
Biological Function |
Pathological Role in CRC |
Research Application |
|
APC |
Negative regulator of Wnt/β-catenin pathway, promotes β-catenin degradation to maintain pathway stability; involved in cell adhesion and migration. |
Germline/somatic mutations lead to loss of function, causing β-catenin accumulation in the nucleus and activating proliferation genes like MYC, a key early driver in carcinogenesis. |
1. Develop APC mutant cell/animal models (e.g., ApcMin/+ mice) to study early carcinogenesis mechanisms. 2. Screen for small molecules that restore APC function. 3. Explore the Wnt pathway's upstream and downstream regulation. |
|
KRAS |
GTPase, serves as a "molecular switch" in EGFR and other pathways; activates downstream MAPK/PI3K pathways, regulating cell proliferation and differentiation. |
Mutations at hotspots (12/13/59/61/117/146) result in loss of GTPase activity, leading to continuous pathway activation, promoting tumour cell proliferation, migration, and drug resistance. |
1. Establish KRAS mutant subtype cell models to analyse functional differences. 2. Test KRAS inhibitors and their effects on downstream molecules. 3. Investigate KRAS interaction with other driver genes. |
|
EGFR |
Transmembrane tyrosine kinase receptor, activates downstream MAPK/PI3K pathways upon ligand binding, regulating cell growth, survival, and angiogenesis. |
Overexpression or pathway dysregulation enhances tumour cell proliferation and invasion; KRAS mutations lead to ligand-independent EGFR activation, contributing to treatment resistance. |
1. Use Co-IP technology to study EGFR protein interactions. 2. Develop EGFR-overexpressing cell models to screen for pathway inhibitors. 3. Investigate the relationship between EGFR expression and other targets. |
|
BRAF V600E |
Serine/threonine kinase, key molecule in MAPK pathway, phosphorylates MEK to activate the pathway, regulating cell proliferation and apoptosis. |
Mutation significantly enhances kinase activity, leading to sustained MAPK activation, accelerating CRC progression and associated with high invasiveness and poor prognosis. |
1. Establish BRAF V600E mutant transplant tumour models to study pathway activation effects on tumour growth. 2. Test the combined effects of BRAF inhibitors and EGFR inhibitors. 3. Explore the link between BRAF mutations and MSI-H phenotype. |
|
VEGF |
Endothelial cell-specific growth factor, promotes endothelial cell proliferation, migration, and angiogenesis upon binding to VEGFR, maintaining tissue blood supply. |
Overexpression in the tumour microenvironment induces angiogenesis, providing nutrients and oxygen to tumour cells, promoting tumour growth and metastasis. |
1. Use endothelial cell models to study VEGF regulation of angiogenesis. 2. Screen for small molecules or siRNAs that inhibit VEGF expression. 3. Analyse the synergy between VEGF and other tumour microenvironment factors (e.g., FGF). |
|
PD-L1 |
Immune checkpoint molecule expressed on immune and tumour cells; binds to PD-1 to suppress T-cell activation and maintain immune tolerance. |
High expression in tumour cells, leading to immune evasion by inhibiting anti-tumour immune responses through PD-1 binding; linked with MSI-H phenotype. |
1. Use immune cell co-culture models to study PD-L1's effect on T-cell function. 2. Screen for transcription factors or signalling molecules regulating PD-L1 expression. 3. Evaluate the effects of anti-PD-L1 antibodies on immune microenvironment remodelling. |
|
HER2 |
Transmembrane tyrosine kinase receptor; dimerises with other HER family members to activate the PI3K/Akt pathway, regulating cell proliferation and survival. |
Gene amplification or overexpression leads to abnormal pathway activation, promoting tumour cell proliferation, migration, and drug resistance, central to HER2-positive CRC subtypes. |
1. Develop HER2-overexpressing cell models to study its effect on cancer cell malignancy. 2. Analyse the cross-regulation between HER2 and EGFR pathways. 3. Test the blocking effect of anti-HER2 antibodies on signalling pathways. |
4. abinScience Recombinant Proteins and Antibodies for Colorectal Cancer Research
|
Category |
Product Name |
Catalogue Number |
Applications |
|
Protein |
Recombinant Human APC Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
|
|
Recombinant Human CTNNB1 Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human GSK3B Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human GSK3B Protein, N-GST |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human TNKS2 Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human KRAS/K-Ras 2 (G12C) Protein, N-GST |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human KRAS Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human EGFR/ERBB1/HER1 Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Rat EGFR/ERBB1/HER1 Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human EGFR Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human BRAF Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human VEGFD Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human VEGFC Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human CD340/ERBB2/HER2/NEU Protein, C-Fc |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Recombinant Human CLDN18/Claudin-18 Protein, N-His |
ELISA, Immunogen, SDS-PAGE, WB, Bioactivity testing in progress |
||
|
Antibody |
Anti-GSK3B Polyclonal Antibody |
ELISA, IHC, WB |
|
|
Anti-CTNNB1 Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Anti-Human CTNNB1 Nanobody |
ELISA, IF, IHC, IP, SPR, WB |
||
|
Anti-TNKS2 Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Anti-KRAS Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Anti-Human KRAS/K-Ras 2 Antibody (SAA1513) |
ELISA |
||
|
Anti-Human EGFR/ERBB1/HER1 Antibody (11F8) |
ELISA, FCM, WB |
||
|
Resaerch Grade Anti-EGFRvIII Antibody (AMG-595) |
ELISA, FACS, Functional assay |
||
|
Anti-Human BRAF/B-Raf (V600E) Antibody (SAA2181) |
ELISA |
||
|
Anti-BRAF Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Anti-VEGFC Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Research Grade Anti-PD-L1 & VEGF Bispecific Antibody (Pm8002) |
ELISA, Bioactivity: FACS, Functional assay, Research in vivo |
||
|
InVivoMAb Anti-Mouse TIGIT & PD-L1 Bispecific Antibody |
FuncS |
||
|
Anti-Human CD274/PD-L1/B7-H1 Antibody (2.7A4) |
Blocking, ELISA, FCM |
||
|
Research Grade Anti-Human CD340/ERBB2/HER2/NEU (MEDI4276) |
ELISA, Bioactivity: FACS, Functional assay, Research in vivo |
||
|
Anti-CD340/ERBB2/HER2/NEU Polyclonal Antibody |
ELISA, IHC, WB |
||
|
Anti-Human CLDN18.2 Antibody (hu7V3) |
ELISA |
AbinScience offers a series of recombinant proteins, antibodies and research biosimilar products related to colorectal cancer research. These cover multiple key targets. For more information, please visit abinScience or email our support team at support@abinscience.com
References
[1] Al-Joufi FA, Setia A, Salem-Bekhit MM, Sahu RK, Alqahtani FY, Widyowati R, Aleanizy FS. Molecular Pathogenesis of Colorectal Cancer with an Emphasis on Recent Advances in Biomarkers, as Well as Nanotechnology-Based Diagnostic and Therapeutic Approaches. Nanomaterials (Basel). 2022 Jan 4;12(1):169.
[2] Bertocchi A, Carloni S, Ravenda PS, Bertalot G, Spadoni I, Lo Cascio A, Gandini S, Lizier M, Braga D, Asnicar F, Segata N, Klaver C, Brescia P, Rossi E, Anselmo A, Guglietta S, Maroli A, Spaggiari P, Tarazona N, Cervantes A, Marsoni S, Lazzari L, Jodice MG, Luise C, Erreni M, Pece S, Di Fiore PP, Viale G, Spinelli A, Pozzi C, Penna G, Rescigno M. Gut vascular barrier impairment leads to intestinal bacteria dissemination and colorectal cancer metastasis to liver. Cancer Cell. 2021 May 10;39(5):708-724.e11.
[3] Tilg H, Adolph TE, Gerner RR, Moschen AR. The Intestinal Microbiota in Colorectal Cancer. Cancer Cell. 2018 Jun 11;33(6):954-964.
[4] Chen Y, Wang D, Li Y, Qi L, Si W, Bo Y, Chen X, Ye Z, Fan H, Liu B, Liu C, Zhang L, Zhang X, Li Z, Zhu L, Wu A, Zhang Z. Spatiotemporal single-cell analysis decodes cellular dynamics underlying different responses to immunotherapy in colorectal cancer. Cancer Cell. 2024 Jul 8;42(7):1268-1285.e7.
[5] Li J, Wu C, Hu H, Qin G, Wu X, Bai F, Zhang J, Cai Y, Huang Y, Wang C, Yang J, Luan Y, Jiang Z, Ling J, Wu Z, Chen Y, Xie Z, Deng Y. Remodeling of the immune and stromal cell compartment by PD-1 blockade in mismatch repair-deficient colorectal cancer. Cancer Cell. 2023 Jun 12;41(6):1152-1169.e7.
[6] Wang X, Fang Y, Liang W, Wong CC, Qin H, Gao Y, Liang M, Song L, Zhang Y, Fan M, Liu C, Lau HC, Xu L, Li X, Song W, Wang J, Wang N, Yang T, Mo M, Zhang X, Fang J, Liao B, Sung JJY, Yu J. Fusobacterium nucleatum facilitates anti-PD-1 therapy in microsatellite stable colorectal cancer. Cancer Cell. 2024 Oct 14;42(10):1729-1746.e8.
[7] Siegel RL, Wagle NS, Cercek A, Smith RA, Jemal A. Colorectal cancer statistics, 2023. CA Cancer J Clin. 2023 May-Jun;73(3):233-254. doi: 10.3322/caac.21772. Epub 2023 Mar 1.
[8] Drew DA, Cao Y, Chan AT. Aspirin and colorectal cancer: the promise of precision chemoprevention. Nat Rev Cancer. 2016 Mar;16(3):173-86. doi: 10.1038/nrc.2016.4. Epub 2016 Feb 12.
[9] Ashktorab H, Brim H. Colorectal cancer subtyping. Nat Rev Cancer. 2022 Feb;22(2):68-69.
[10] Dienstmann R, Vermeulen L, Guinney J, Kopetz S, Tejpar S, Tabernero J. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer. 2017 Feb;17(2):79-92. doi: 10.1038/nrc.2016.126. Epub 2017 Jan 4. Erratum in: Nat Rev Cancer. 2017 Mar 23;17(4):268.
+86-027-65523339
중국 우한시 심둔사로 666번지 C동, 우한, 430206

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский