Breast cancer is the most common malignancy in women worldwide, accounting for approximately 11.6% of new cancer cases and 6.9% of cancer deaths, posing a serious threat to women's health. Although early diagnosis and treatment have improved significantly, many patients still experience disease progression and distant metastasis, and the underlying molecular mechanisms remain incompletely understood. Circular RNAs (circRNAs) are a class of non-coding RNAs with covalently closed loop structures that exert diverse biological functions in tumorigenesis, including acting as miRNA sponges, binding RNA-binding proteins, and even encoding peptides, making them a hotspot in cancer research. Tumor-associated macrophages (TAMs), a key component of the tumor microenvironment, engage in critical “cancer-immune cell” crosstalk with breast cancer cells and play an essential role in tumor progression.
Recently, a research team from China Agricultural University published a study in Oncogene that comprehensively elucidates the role and molecular mechanism of circCLASP1 in breast cancer, providing new targets and insights for diagnosis and therapy.
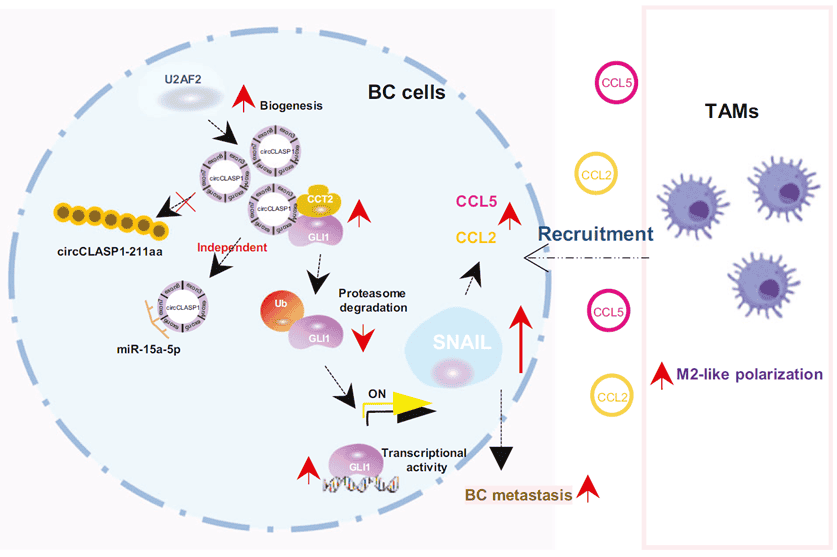
Figure 1. Graphical abstract
circCLASP1 is highly expressed in breast cancer tissues and serum and shows diagnostic potential
By re-analyzing the GEO dataset (GSE71651), the team identified circRNAs significantly differentially expressed in breast cancer and found that circCLASP1 was one of the most strongly upregulated molecules. Validation in 65 breast cancer tissues, 42 adjacent normal tissues, serum from 61 patients, and 10 healthy controls confirmed markedly elevated circCLASP1 levels in both tumor tissues and patient serum. Its expression positively correlated with lymph node metastasis, Ki67 status, and tumor size. circCLASP1 exhibited a longer half-life than linear CLASP1 mRNA, resisted RNase R digestion, and was predominantly localized in the cytoplasm. These findings suggest that circCLASP1 holds promise as a diagnostic biomarker for breast cancer.
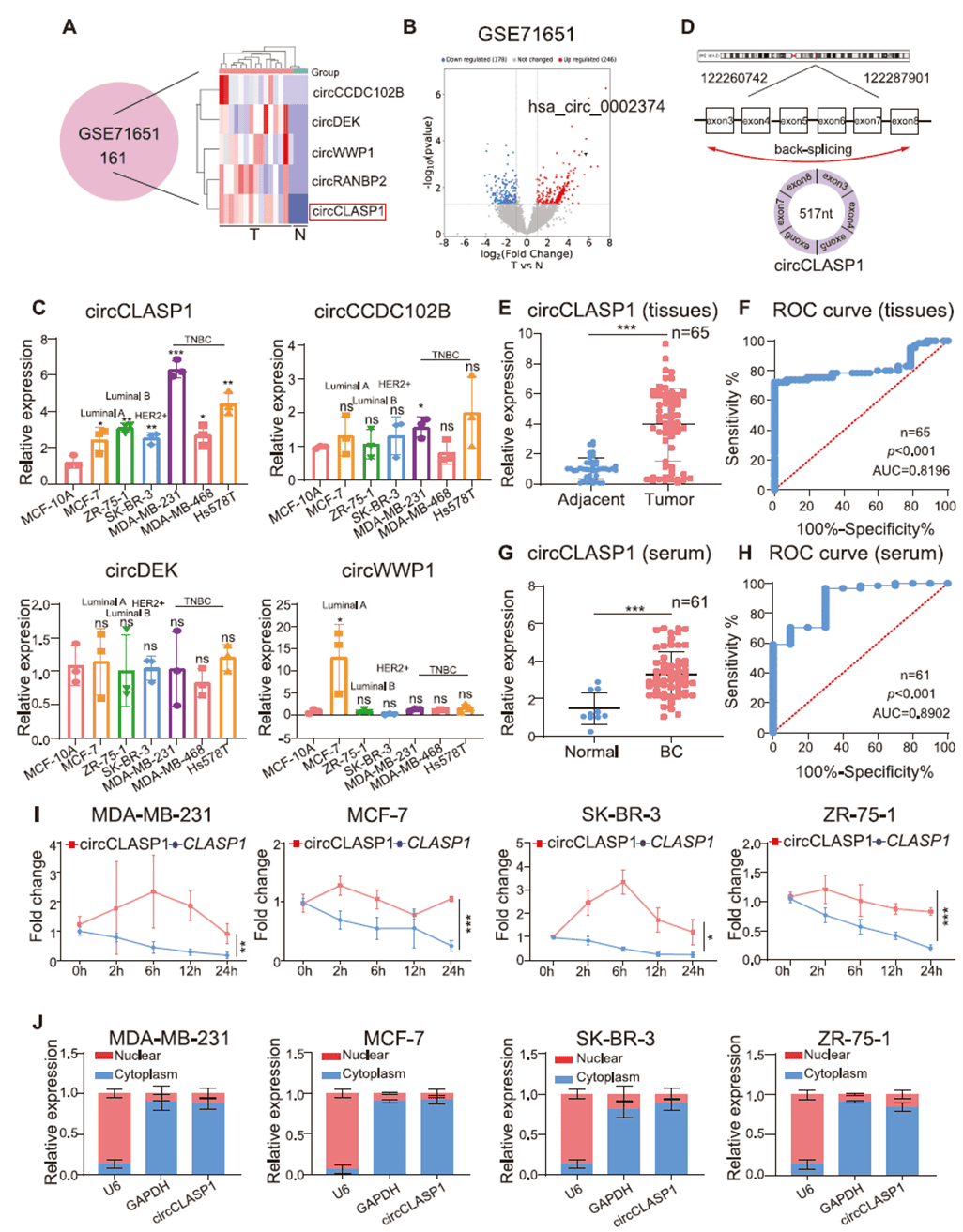
Figure 2. Identification and characterization of circCLASP1 in breast cancer
circCLASP1 promotes proliferation, migration, and invasion of breast cancer cells
Using shRNAs targeting the backsplice junction, the researchers successfully knocked down circCLASP1 in breast cancer cell lines. Loss-of-function experiments showed that silencing circCLASP1 significantly impaired proliferation, colony formation, and Ki67 positivity in MDA-MB-231 and MCF-7 cells, and reduced growth and size of patient-derived organoids (PDOs). Transwell assays demonstrated suppressed migration and invasion. Cell-cycle analysis revealed G0/G1 arrest and increased apoptosis, with elevated BAX and reduced BCL2 levels. In vivo, circCLASP1-knockdown tumors in nude mice were significantly smaller. Collectively, these data establish circCLASP1 as an oncogenic driver that promotes proliferation, migration, and invasion in breast cancer.
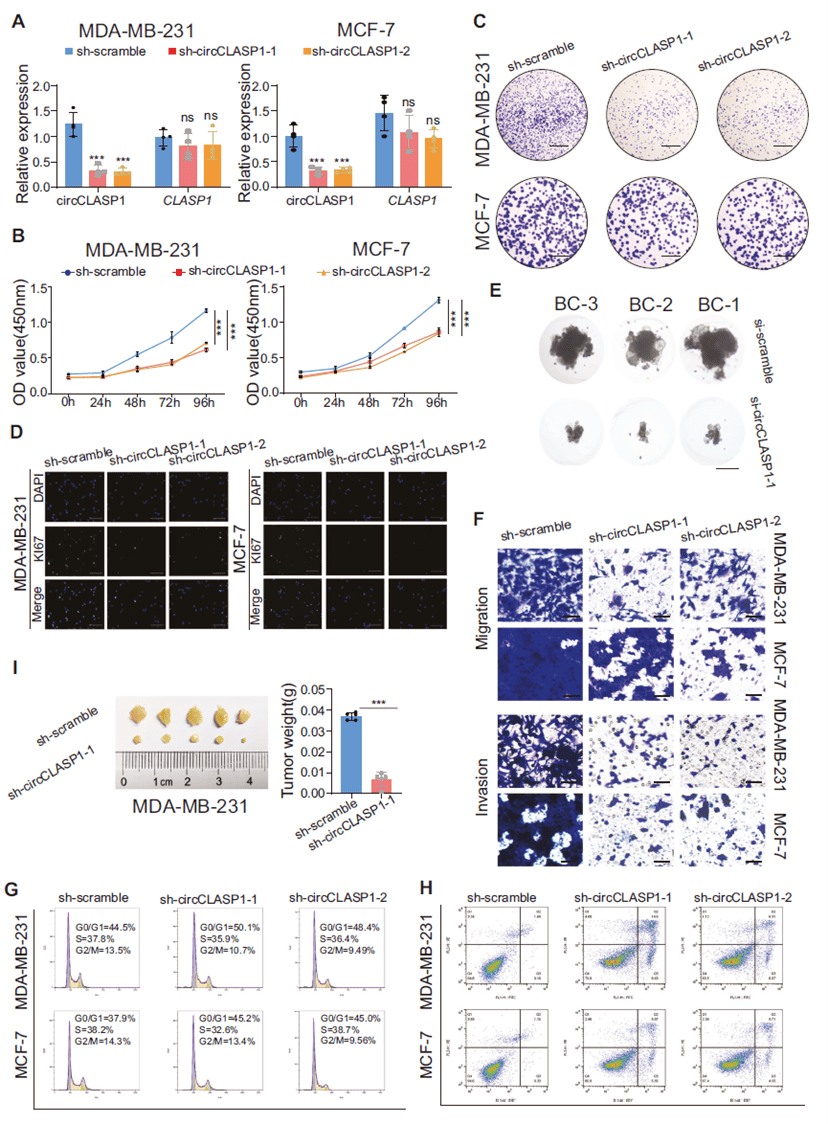
Figure 3. circCLASP1 functions as an oncogene in breast cancer cells
U2AF2 regulates circCLASP1 biogenesis and contributes to malignant phenotypes
RNA-binding proteins (RBPs) can modulate circRNA formation by binding flanking intronic sequences. By integrating three RBP prediction databases, the team identified PTBP1 and U2AF2 as candidate upstream regulators. Overexpression experiments showed that only U2AF2 significantly increased circCLASP1 levels. U2AF2 was highly expressed in breast cancer tissues, correlated with poor overall survival in lymph-node-positive and Ki67-high patients, and positively correlated with circCLASP1 expression. RIP assays confirmed U2AF2 binds the “+112~+136” region in the flanking sequence to promote backsplicing. Rescue experiments demonstrated that U2AF2 overexpression enhanced proliferation, migration, and invasion, effects partially reversed by circCLASP1 knockdown.
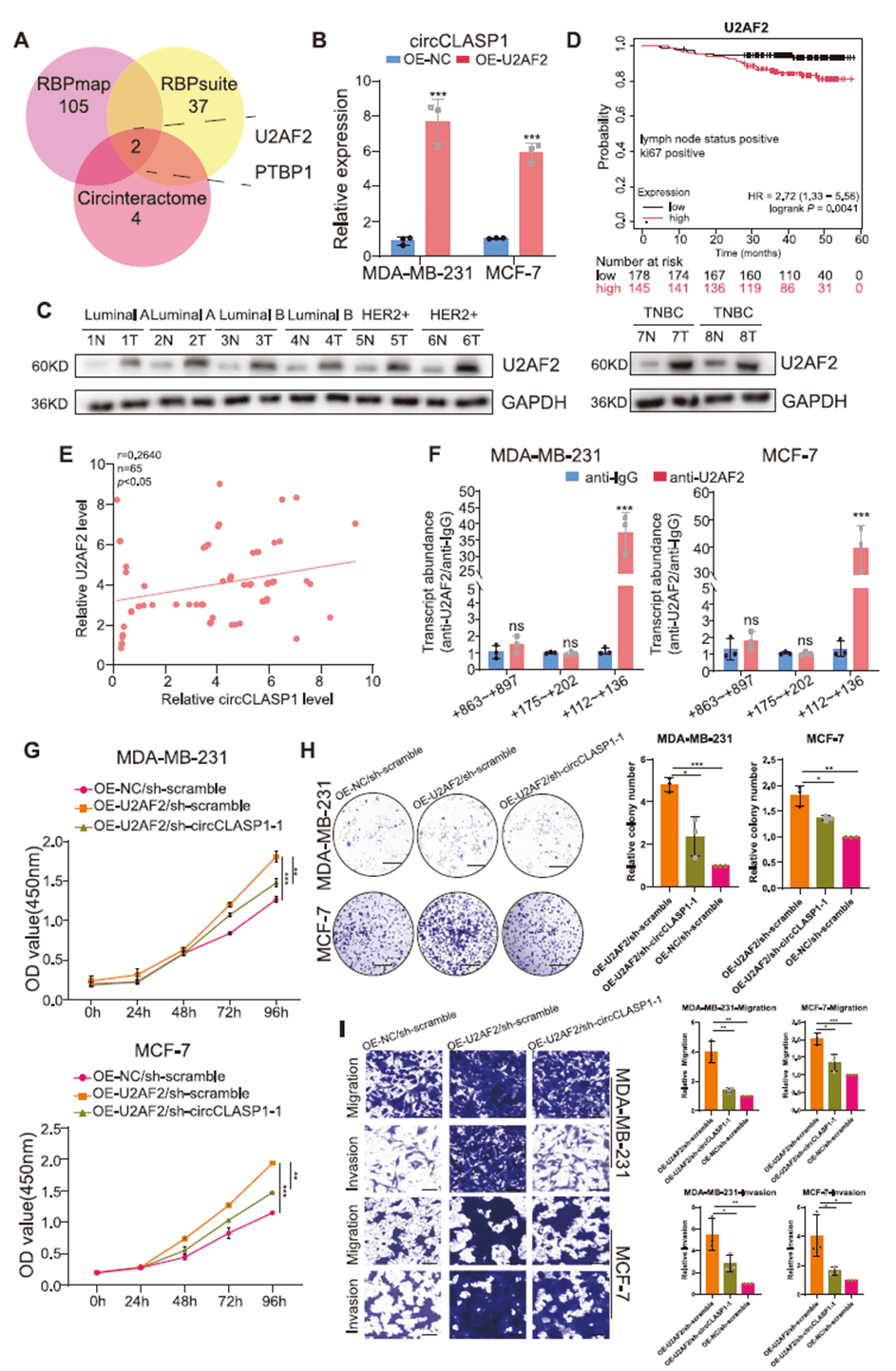
Figure 4. U2AF2 mediates the biogenesis of circCLASP1
circCLASP1 does not function via miRNA sponging or peptide encoding
Although cytoplasmic and able to bind AGO2, circCLASP1 was predicted and validated to directly interact with miR-15a-5p. However, overexpressing miR-15a-5p failed to reverse the oncogenic effects of circCLASP1 overexpression, indicating that circCLASP1 does not exert its oncogenic role through miRNA sponging or peptide translation.
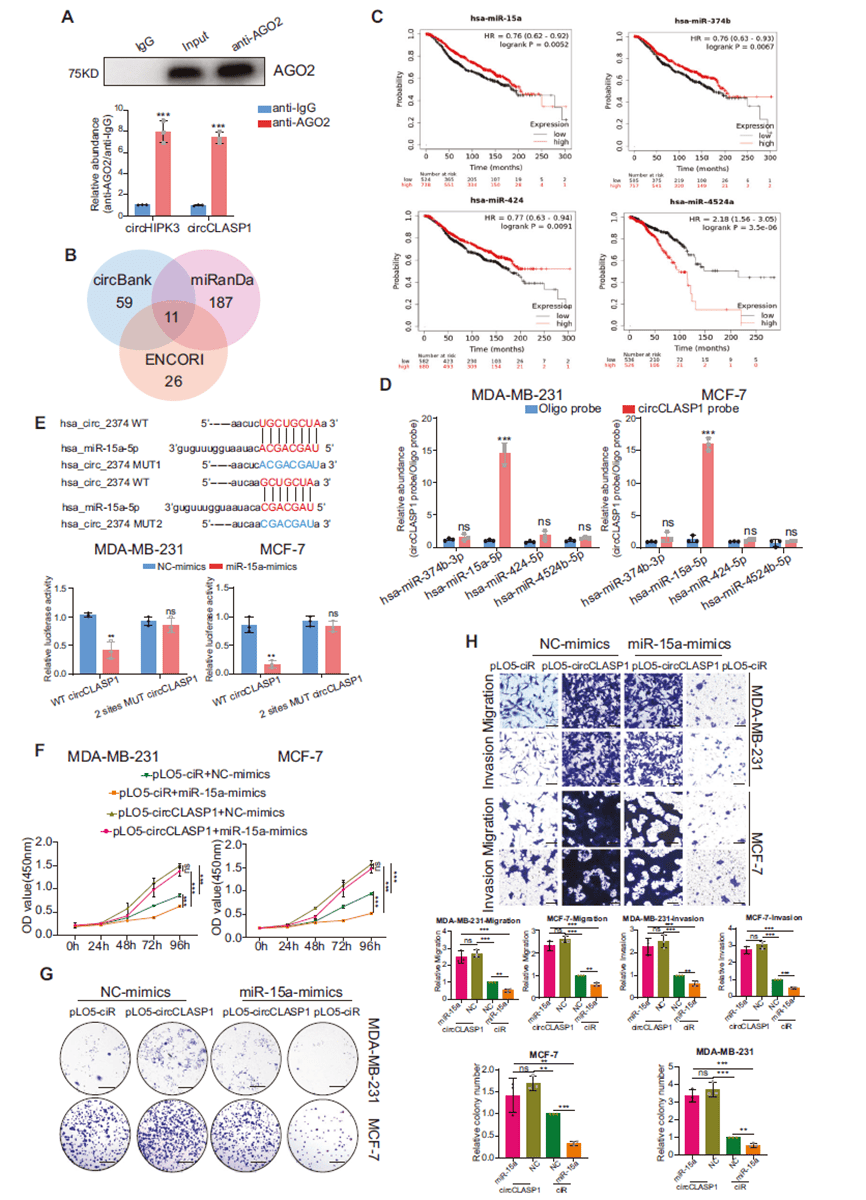
Figure 5. circCLASP1 promotes breast cancer progression without acting as a miRNA sponge
circCLASP1 binds CCT2 and inhibits ubiquitin-mediated degradation of GLI1
RNA pull-down followed by mass spectrometry identified the chaperonin CCT2 as a specific interacting protein. The interaction was validated by RIP, and AlphaFold3 predicted a stable binding interface. Co-IP showed that circCLASP1 enhances the CCT2–GLI1 interaction, forming a ternary complex. Knockdown of circCLASP1 reduced GLI1 protein stability and half-life without affecting mRNA levels, an effect rescued by the proteasome inhibitor MG132. Ubiquitination assays confirmed increased GLI1 ubiquitination upon circCLASP1 depletion. (Anti-CDK4 antibody used in this study was provided by AtaGenix.) Thus, circCLASP1 stabilizes GLI1 protein by binding CCT2 and preventing its ubiquitin-mediated degradation.
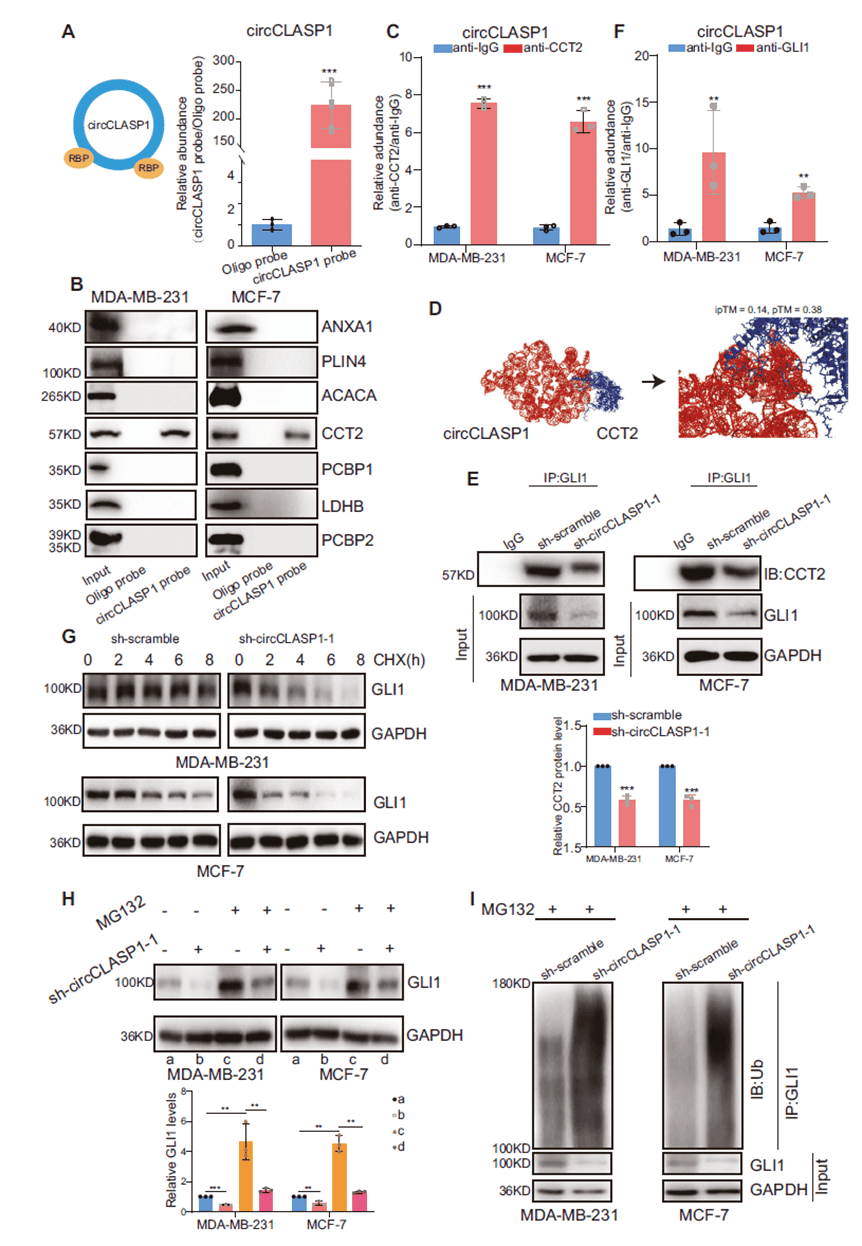
Figure 6. circCLASP1 binds CCT2 and modulates ubiquitin-mediated GLI1 degradation
circCLASP1 drives breast cancer progression via the CCT2/GLI1/SNAIL axis
circCLASP1 knockdown reduced nuclear accumulation of GLI1. High SNAIL expression (a downstream target of GLI1) was associated with poorer survival, and SNAIL levels positively correlated with both GLI1 and circCLASP1 in clinical samples. Overexpression of either wild-type or miR-15a-5p-binding-site-mutated circCLASP1 similarly upregulated SNAIL and promoted malignant phenotypes, confirming that the CCT2/GLI1/SNAIL axis is the primary mechanism and is independent of miR-15a-5p.
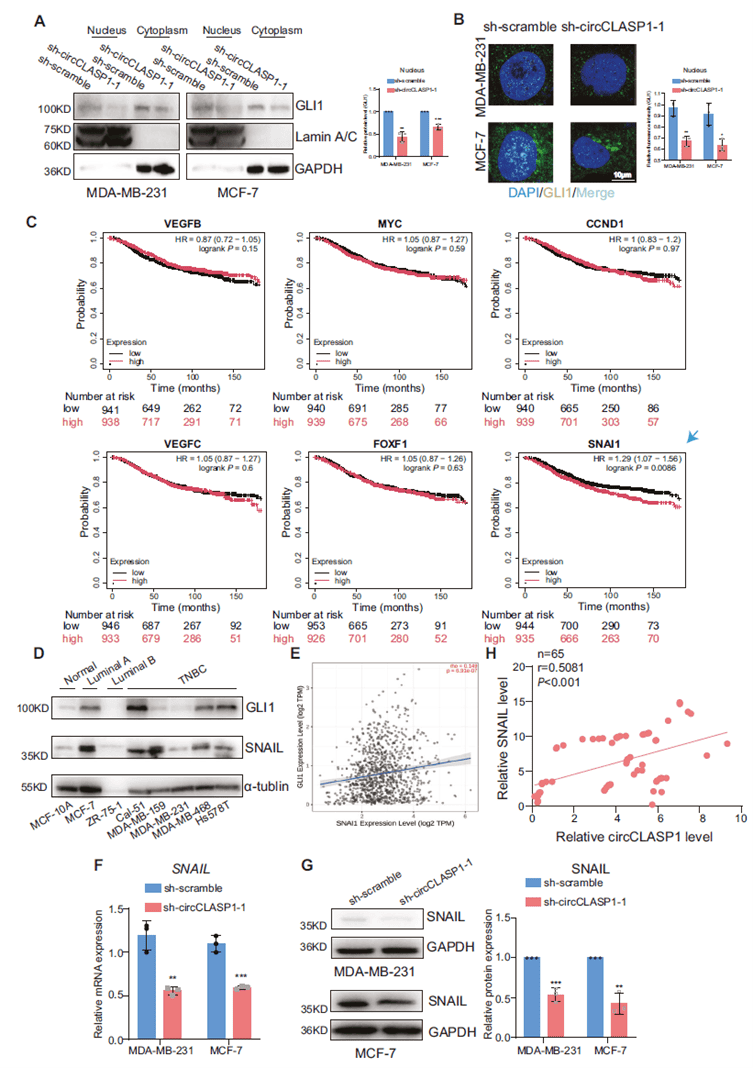
Figure 7. circCLASP1 regulates SNAIL expression via GLI1 at the transcriptional level
circCLASP1 promotes macrophage recruitment and M2 polarization through SNAIL, accelerating lung metastasis
IHC showed fewer CD68+ macrophages (but not CD3+ T cells) in tumors with low circCLASP1. Conditioned medium from circCLASP1-overexpressing cells drove THP-1 monocytes toward an M2 phenotype, while knockdown had the opposite effect. circCLASP1 upregulated CCL2 and CCL5 expression in a SNAIL-dependent manner, enhancing macrophage recruitment in vitro and in vivo. In a 4T1 tail-vein lung metastasis model, circCLASP1 overexpression dramatically increased metastatic nodules, Snail expression, and macrophage infiltration—effects abrogated by Snail knockdown. Thus, circCLASP1 accelerates breast cancer metastasis by remodeling the tumor immune microenvironment through the SNAIL–CCL2/CCL5 axis.
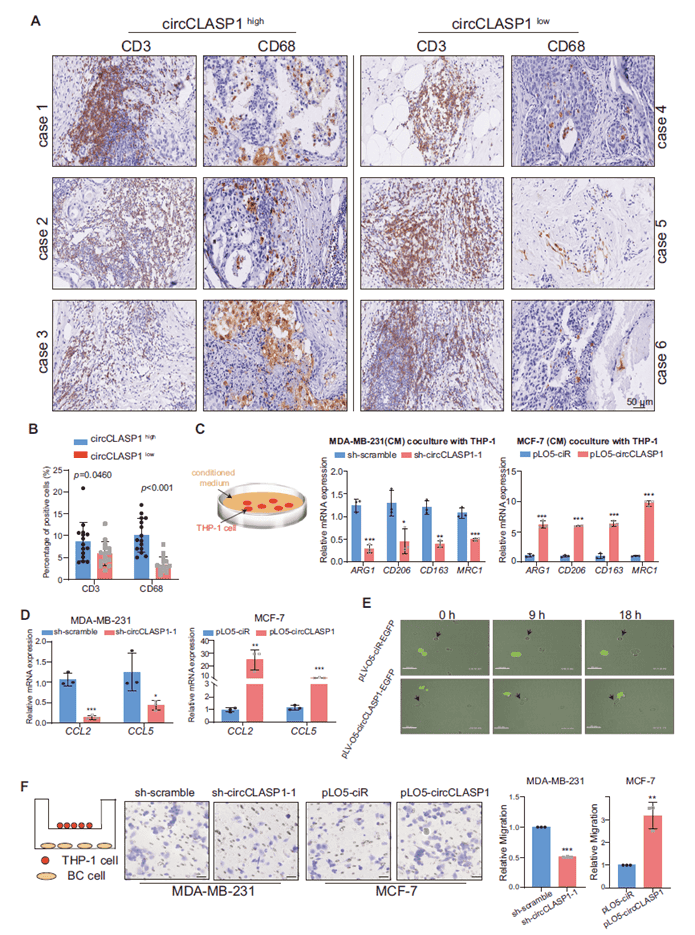
Figure 8. circCLASP1 enhances macrophage recruitment in breast cancer cells
This study systematically demonstrates that circCLASP1 is highly expressed in breast cancer tissues and serum, correlates with adverse clinicopathological features, and possesses diagnostic biomarker potential. Mechanistically, circCLASP1 is generated through U2AF2-mediated backsplicing, acts as a scaffold to bind CCT2, stabilizes GLI1 by blocking its ubiquitination and degradation, drives nuclear GLI1 accumulation, upregulates SNAIL, and ultimately recruits and polarizes M2 macrophages via CCL2/CCL5, thereby fueling proliferation, invasion, and lung metastasis. These findings uncover a critical oncogenic network centered on circCLASP1 and provide novel diagnostic and therapeutic targets for breast cancer.
abinScience was founded in Strasbourg, France, leveraging the region’s outstanding research ecosystem to develop and produce high-quality life science reagents. Guided by the vision “Empowering Bioscience Discovery,” abinScience is committed to providing reliable, efficient experimental solutions to researchers worldwide.
abinScience provided the anti-CDK4 antibody used in this study, ensuring accurate protein detection and supporting robust validation of the circCLASP1–CCT2–GLI1–SNAIL signaling cascade.
Disclaimer: This article is a summary based on publicly available literature. All products mentioned are for research use only.

 中文
中文 English
English 한국어
한국어 日本語
日本語 Español
Español Français
Français Русский
Русский










